Transforming high-plex images into biological insights doesn’t have to be difficult. A new collaboration provides a powerful and efficient pipeline from instrument to data analysis.
The field of spatial biology is continuing to grow at a rapid pace – and that trend is likely to continue. With spatial technologies becoming a standard in translational research, it’s crucial for researchers to have the right tools to clearly see the ‘how and where’ of cellular interactions with accuracy. In this new wave of information, it’s becoming more critical than ever to adopt end-to-end solutions that take the complexity out of spatial imaging.
The second session of our current IMC Forum series on data analysis saw James Mansfield, Senior Vice President of Research Business Development at Visiopharm®, introduce Phenoplex™, an analysis software that has been paired with Standard BioTools™ Hyperion™ and Hyperion+ Imaging Systems to provide a streamlined sample-to-analysis workflow solution for high-plex data.
Phenoplex provides a complete workflow for multiplex image analysis in seven steps:
Here, we share a quick overview of each step and how they can help you tailor your analysis to your biological question.

Phenoplex provides a complete workflow for multiplex image analysis in seven steps:
Phenoplex Workflow
Visualize
- Read all major multiplex file formats, including native MCD files
- Convert MCD files into pyramidal TIFFs
- Set up panels of colors for easy switching between biologically relevant groupings of markers
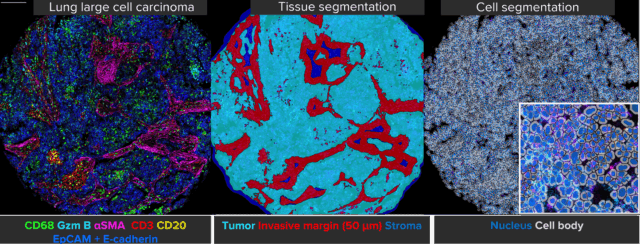
Classify Tissues
- Utilize paint-to-train deep-learning-based tissue segmentation
- Remove artifacts
- Compartmentalize tissue into regions and get boundary information
Detect Cells
- Detect cells using AI algorithms, including one specifically designed for IMC™ Ir nuclear stains
- Augment algorithms using user data and cell-type annotations
Phenotype
- Choose an automated method that uses an unsupervised clustering algorithm to find the most common marker combinations
- Or opt for a guided method that helps you set gates/thresholds for optimum positivity settings
Verify
- Solve the problem of determining whether cells have the correct phenotype
- Use interactive tools to check biomarker expression and cell phenotypes and expression patterns
Explore
- Utilize a set of interactive plotting tools to explore your data and review whether phenotypes are correct
- Investigate distances between boundaries and phenotypes, apply analysis tools such as neighborhood analysis and classify cells based on their tissue compartment
Publish
- Export your data as a multitude of different industry-standard file types
Transforming high-plex images into biological insights doesn’t have to be difficult. This collaboration provides a powerful and efficient pipeline from instrument to data analysis, making it easier to arrive at confident decisions impacting healthcare.
Click below to watch Mansfield’s talk to learn about the specific ways Phenoplex can facilitate your image analysis, along with hands-on examples of defining cell types and understanding tissue architecture.
Images courtesy of James Mansfield
Unless explicitly and expressly stated otherwise, all products are provided for Research Use Only, not for use in diagnostic procedures. Find more information here.