Watch this talk from Jennifer Gorman to learn how pushing the boundaries of spatial proteomics can save time and lead to more in-depth observations of heterogeneity
Jennifer Gorman, PhD, is a Scientific Associate and Lab Manager at the Lunenfeld-Tanenbaum Research Institute in Toronto, working with Hart Jackson, PhD, to push the boundaries of the field of high-plex imaging using Imaging Mass Cytometry™ technology. The Jackson Lab has focused on scaling – looking at more tissues, more tissue area and more samples – to better capture and ultimately understand the heterogeneity that makes up disease complexity.
Advances in IMC™ technology are helping researchers achieve this. The use of whole slide imaging modes enables in-depth exploration of tissue heterogeneity:
- Preview Mode allows for rapid, simultaneous 40-plus-marker visualization across an entire tissue section, facilitating the identification of regions of interest for use in Cell Mode analysis
- Cell Mode and downstream single-cell analysis provide detailed insights at the highest resolution, uncovering distinct neuronal populations and their protein expression profiles, which is crucial for understanding the mechanisms of disease
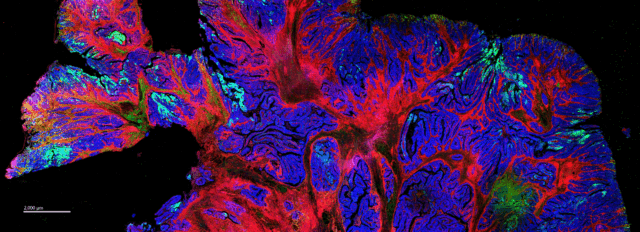
- Tissue Mode enables comprehensive visualization of all markers, revealing the heterogeneity of aggregate distribution across the whole tissue
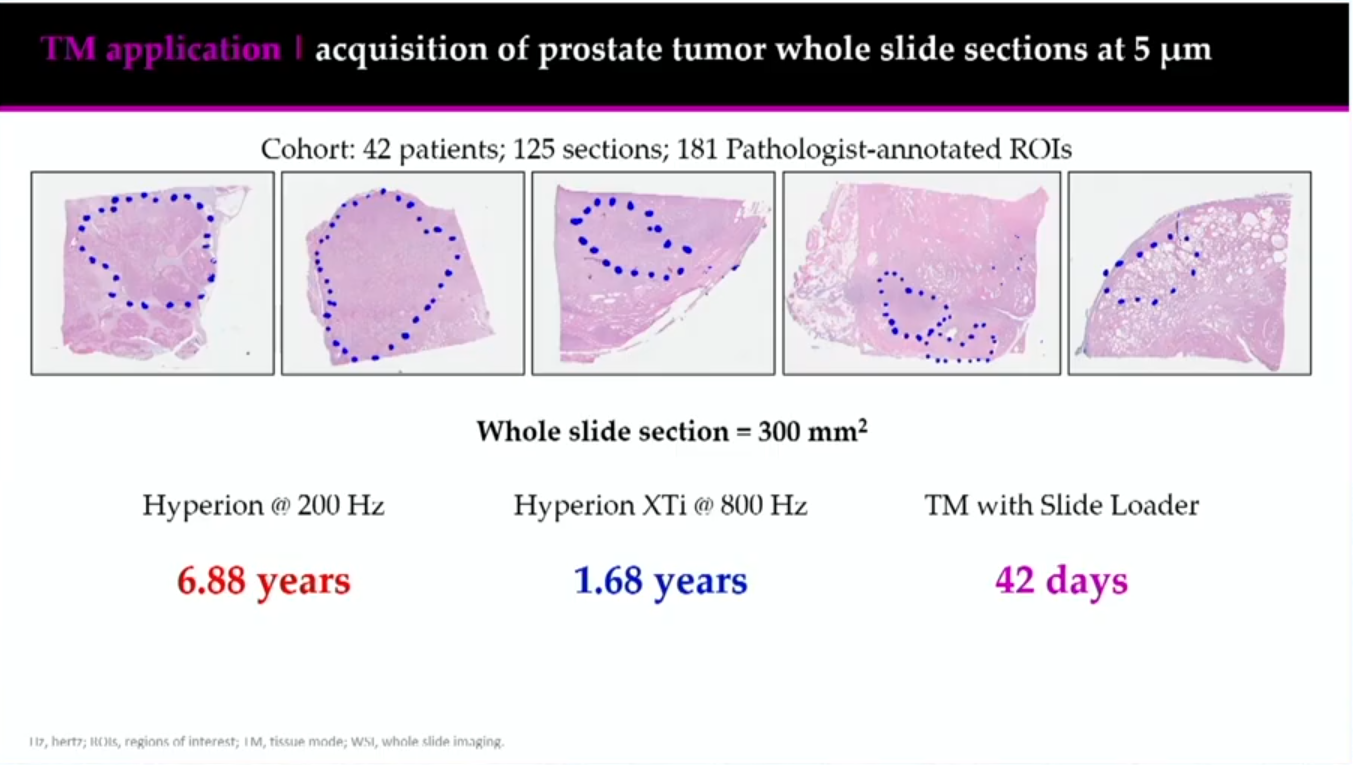
Gorman and the team applied a high-throughput Tissue Mode to investigate the proteomic characterization of high-risk prostate cancer, with the aim of assessing differences between Gleason pattern 4 and Gleason pattern 5 histologies and examining alterations in tumor vs. non-malignant tissue.
The cohort was large: 125 sections and 181 pathologist-annotated regions of interest (ROIs) from 42 participants. The team wanted to determine the best, most efficient way to image this large cohort. They determined acquisition times for 1 µm and 5 µm resolution imaging and what area of the slides they could image at each resolution, ultimately deciding to raster the data at 5 µm using Tissue Mode and image the entire slide.
This faster acquisition time opened up a new avenue for Gorman. “We could do 4,500 by 4,000 µm, or we could everything,” she says. “And so the decision was made that we would actually do this on the entire tissue section at 5 µm” as opposed to 1 µm.
She says that changing the rasterization size from 1 µm to 5 µm using Tissue Mode allowed them to go 25x faster than before. In addition, the slide loader allowed Gorman and her team to run samples on the Hyperion XTi Imaging System overnight or during the weekend without having someone there to physically change the slides, leading to an even greater time savings.
Acquisition at 5 µm allowed for a better look at spatial heterogeneity not just in the pathologist-annotated regions but across the entire tissue. In addition to immune infiltration, they also observed alterations in metabolic and signaling markers across samples: “This was really important for us, because had we actually done this acquisition at 1 µm, we never would have noticed that,” Gorman says.
What’s next for the lab, and how does this information get analyzed? “We’re taking different approaches and seeing what we can adapt to work with this 5 µm data,” Gorman says; she and her team are applying spot-counting and pixel-based methods to quantify and cluster the data.